To work in Abigail Salyers’ laboratory at the University of Illinois, is to play matchmaker to some unlikely couples. Standing at her laboratory bench, PhD student Kaja Malanowska lifts the cover from a petri dish to pick up a half a billion or so Escherichia coli bacteria with the tip of a sterile needle. She adds them to a tube containing about as many Bacteroides thetaiotaomicron floating in a tiny puddle of antibiotic-laced broth. Finally, Malanowska transfers the tube to an incubator set at 98.6° F – human body temperature – where she will leave them to co-mingle for the night.
What Malanowska will find in the morning is no secret to her mentor. “In the bacterial world’s version of casual sex, E. coli and B. theta will swap a few genes,” explains Salyers
For decades, microbiologists have known that bacteria employ a variety of methods to share genes between species, with each gene or group of genes spelling out the DNA instructions for a potentially useful trait. What initially took Salyers by surprise was to discover how indiscriminate the pairings – with heavily armored “gram-negative” organisms such as E. coli sharing genes with near-naked “gram-positive” bacteria at the opposite end of the bacterial kingdom.
“It’s rather like an armadillo mating with a squid,” she muses.
Three things make this finding far more than a biological curiosity. One: B. theta and E. coli number among the most abundant of the trillions of bacteria that live in the human colon. Two: The DNA they’ll be swapping in Malanowska’s test tubes tonight will be genes for antibiotic resistance. And three: This kind of dangerous gene exchange may be as common inside people as it is in Slayer’s laboratory incubator.
What’s more, Salyers knows that the antibiotic in her student’s test tube will not simply fail to kill the bacteria floating inside it. It will act as an aphrodisiac.
“Tetracycline really turns these guys on,” says Salyers. Indeed, exposure to various antibiotics goads many, if not most, known bacteria to begin copying and swapping “survival” genes, including genes for drug resistance.
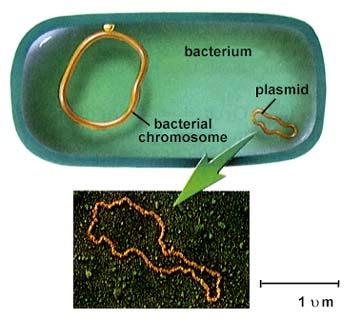
Salyers team has found that an overnight tête-à-tête is more than enough for these simple, one-celled organisms to complete their intimate exchanges, which they accomplish by copying and transferring snippets and circlets of auxiliary genes called transposons and plasmids. Indeed, Salyers has documented the transformations wrapping up in as little as an hour.
Clearly, such goings-on have disturbing implications. As most everyone knows from headlines and TV exposes, antibiotic resistance has become the scourge of 21st-century medicine. In fact, in the 60 years since the first antibiotics came into widespread use, we’ve bred varying degrees of drug resistance into every known type of disease-causing bacterium. Especially scary has been the recent spread of multi-drug-resistant strains of staph and strep bacteria, not only in hospitals but also through entire communities, with increasingly deadly results.
What has largely escaped notice in the hubbub over drug-resistant disease is that antibiotics don’t just breed resistance into the infection-causing “bad guys” they’re designed to kill.
“Every time your physician treats you for an infection, that physician is also treating the bacteria that normally reside in your intestines, on your skin, and throughout your nose, mouth and respiratory tract,” Salyers explains. The result has been a steep increase in drug-resistance in these resident, or commensal, bacteria, including Bacteroides, which makes up a quarter to a third of the rain-forest rich community of micro-organisms living in the human gastrointestinal tract.
Though normally harmless, Bacteroides and many of its neighbors can cause life-threatening infections if they enter an open wound or surgical incision. “A person who has antibiotic-resistant Bacteroides is a ticking time bomb,” Salyers warns. More disturbing, perhaps, is the realization that over the last half century we’ve turned our intestinal “microflora” into reservoirs of drug-resistance genes that can be passed to the pathogens, or “professional” disease-causing bacteria, that regularly pass through our digestive tract on contaminated food as well as the inhaled and swallowed secretions from a throat infection (or anything else we stick in our mouths).
Indeed, Salyers and others have used DNA fingerprinting to build a strong case that B. theta picked up one or more of its resistance genes from tetracycline-resistant strains of strep and staph. Before 1970, less than a third of Bacteroides sampled from the colons of healthy people contained genes for resistance to tetracycline drugs, Salyers found. By the late 1990s, the gene was showing up in over 80 percent of Slayer’s samples, and more than 15 percent of those had picked up one to three additional genes for resistance to erythromycins, another large and important class of antibiotics.
Other studies have found similar trends in other common intestinal bacteria. A recent Harvard study of incoming hospital patients, for example, found that close to 20 percent of patients harbored E. coli resistant to three or more groups of antibiotics. Fifteen percent harbored drug-resistant strains of the equally common intestinal bacteria Enterobacter, Pseudomonas, and Klebsiella.Scariest of all, perhaps, 10 percent of these bacteria proved resistant to five different groups of antibiotics – virtually everything in a doctor’s standard arsenal.
Most worrisome of all, are bacterial genes that convey resistance to vancomycin – one of the last drugs still effective against otherwise unstoppable strains of staph and strep. Over the last two decades, hospitals have become particular hotbeds for vancomycin-resistant Enterococcus (VRE). Like B. theta, Enterococcus is an intestinal microbe that does not normally cause problems. But it can cause life-threatening sepsis, or blood infection, when it enters the bloodstream via contaminated medical devices such as catheters, IV lines, and surgical implants.
Scariest of all is what can happen when VRE meets MRSA, or methicillin-resistant Staph aureus. Just such a coupling has been documented at three times in North American. The damnable progeny: VRSA. That is, a staph infection resistant to all standard antibiotics.
Adding to the bad news have been drug-resistant outbreaks of the toxin-producing bacterium Clostridium difficile. A common member of the intestine’s microbial ecosystem, C. difficile is normally kept in check by more abundant and better-behaved bacteria such as Bacteroides. Problems tend to arise when someone takes antibiotics – especially broad-spectrum antibiotics – which can raze the intestinal neighborhood of everything but…you guessed it, C. difficile. C. difficile survives antibiotic assaults by retreating into a dormant spore form. Once the drugs have cleared the body, it can spring back to life and overgrow before other bacteria have a chance to move back in.
Because of the toxins C. difficile produces, this overgrowth can cause a range of problems, from what’s commonly known as “antibiotic-associated diarrhea” to painful colitis and even deadly perforations of the intestines. The latter scenario remained mercifully rare until recently, with the development of several new strains of C. difficile with drug-resistance genes that obviates their need to retreat into spore form in the face of antibiotics. In other words, they can begin overgrowing in a patient’s intestines as soon as he or she starts taking antibiotics.
Worst of all, some of these strains have a mutation in the gene that normally limits the organism’s toxin production. And in the last three years, these super-virulent bugs have killed hundreds of hospital patients in the U.S., Canada, and the United Kingdom. Admittedly, some of the victims were already gravely ill. But some had checked into the hospital for relatively minor surgical procedures.
All this said, the body’s intestinal bacteria remain far more beneficial than dangerous. We now know that they serve vital functions. They digest foods that our own bodies can’t and provide us with added nutrition, including vitamins, in the process. They also prime the body’s disease-fighting immune system to be on the lookout for truly dangerous microbes. New research even suggests that a normal and undisturbed complement of intestinal bacteria plays a vital role in restraining our immune system from over-reacting to harmless substances such as the pollen and dander that trigger allergies and asthma. And like the normal, resident bacteria of the skin, mouth, nose, and genital tract, those of the intestine form a kind of living mulch that prevents less-well-adapted microbes from getting established.
So how do medical experts propose to handle the fact that we’ve now imbued the body’s ever-present microflora with dangerous genes for drug resistance?
In the short term, the desperate cry is for newer, more powerful antibiotics. In 2004, the physician-led Infectious Disease Society of America launched its “Bad Bugs, No Drugs” campaign for increased government funding and industry interest in re-invigorating the war on drug-resistant microbes.
“The pipeline for antibiotics has been drying up,” IDSA president Walter Stamm warned. “We need the pharmaceutical industry to get back in the fight before it’s too late.” Backing up such concerns, studies show that large pharmaceutical companies have been significantly slowing, even abandoning, antibiotic research in favor of more profitable drugs for chronic conditions such as heart disease and arthritis.
At the same time, who could blame the industry from shying away from investing in drugs that might remain effective for less time and money than it took to develop them. On average, bacteria develop resistance to a new antibiotic within three years of the drug’s widespread introduction. And even the most strident advocates of antibiotic research no longer talk of “defeating” the enemy, but simply keeping up with it. With dark humor, our arms race with infectious disease has been likened to the conundrum that Alice found herself in after stepping through the Looking Glass. Or in the words of the Red Queen: “Here, you see, it takes all the running you can do to keep in the same place.”
Unfortunately, fighting drug resistance with “more of the same” only worsens the inherent problems of breeding resistance into the body’s “good germs,” as well as those that imbue our food, soil, and water – all of which have become increasingly contaminated with antibiotics.
“The antibiotic paradox is that these drugs will always sow the seeds of their own downfall by selecting for bacteria that can resist their activity,” explains Tufts University microbiologist Stuart Levy. In 1981, Levy founded the Alliance for Prudent Use of Antibiotics, an international organization committed to slowing and preventing the development of drug-resistance by reining in the overuse and misuse of these miracle drugs.
The U.S. Centers for Disease Control joined the effort in 2004 with its own “12-Step Program” for reducing resistance by breaking physicians’ antibiotic “addiction.” Specifically, the CDC campaign aims to break problematic habits such as doctors using antibiotics prophylactically – that is, as a hedge against possible infections – and their keeping patients on antibiotics too long – that is, after an infection has cleared.
Of particular concern to Levy, Salyers, and others studying resistance in commensal bacteria is the long-term use of low-dose antibiotics such as tetracycline for acne-prune teenagers and amoxicillin to prevent chronic ear infections in babies and preschoolers.
It also makes a huge difference what kind of antibiotics a doctor chooses, says infectious-disease specialist Curtis Donskey, of Case Western University, in Cleveland. “Some antibiotics disturb the body’s microbiota more than others,” he explains. Worst of all may be the broad-spectrum, or “big gun” antibiotics that doctors often prescribe when they’re not sure of what they’re treating. Except in immediately life-threatening situations, says Donskey, these doctors need to be running tests to pinpoint which, if any, bacterium is causing an illness and then target it with a narrow-spectrum “bullet” of an antibiotic, less likely to affect the body’s bacterial bystanders.
Yet Levy, Salyers, and others warn that doctor prescriptions may represent but a fraction of the antibiotics we meet each day. They point to the tons of antibiotic drugs routinely added to livestock feed to boost growth and prevent the spread of infections among animals crowded into factory-style warehouses and pens. Some of these drugs end up in surrounding water supplies, as well as milk and meat – though it remains unclear whether these trace amounts are enough to breed resistance.
More worrisome, says Salyers, “there’s now no question that feeding all these antibiotics to livestock selects for resistant bacteria in their intestines and these then end up in the meat on the supermarket shelf”…and just a fork away from co-mingling with our own bacterial nation within. In fact, DNA fingerprinting of various resistance genes has shown that they have moved between the resident bacteria of pigs, cows, and those humans. “But we’re like archaeologists when it comes to following past transfers,” Salyers adds. “It’s usually impossible to determine the direction.” That is, whose microbes transferred their drug-resistance genes to whom?
The good news is that solutions, at least partial solutions, exist to many of the above problems. In 1998, the nations of the European Union banned the use of antibiotics as livestock growth enhancers and restricted which classes of antibiotics the industry could use to treat infections in food animals. Subsequent studies have shown that the rule change has slowed, and in some cases partially reversed, the development of drug resistance in animals, food, and possibly patients. At present, however, North American lawmakers, facing strong lobbying by the livestock industry, have declined to follow suit, at least without stronger proof of harm to human health.
Meanwhile, doctors appear to have reined in their prescribing practices somewhat, says Donskey. “Still, we’re finding that about a third of antibiotics being taken in this country prove unnecessary,” he adds. But such gains may be lost with a keen, new interest in using antibiotics to treat a wide range of chronic inflammatory diseases such as multiple sclerosis, rheumatoid arthritis, even such widespread ills as atherosclerosis. Growing evidence suggests that the chronic inflammation behind these disorders may, in fact, result when a person’s immune system over-reacts to trace amounts of bacteria in body tissues such as joints and arteries. However, in the rush to treat with antibiotics, proponents have not yet factored in the possible harm.
“Unfortunately, most physicians still think of antibiotics as something benign,” says Donskey. “And that has to stop.”
Of course, even careful and appropriate use of antibiotics still breeds resistance, albeit more slowly. One way to reduce the effect on the body’s resident microbes may lie in experimental compounds that de-activate antibiotics before they reach the teeming bacterial community of the large intestine. Tests in animals show that the de-activation does not reduce antibiotic effectiveness because the drugs pass into the bloodstream higher up in the digestive tract.
Others are exploring ways to use good bacteria to fight the bad. Scandinavian pediatricians, for example, have had success stopping chronic ear infections by immediately following antibiotic treatment with nasal sprays containing bacteria cultured from the noses of healthy children. Presumably, these organisms colonize a toddler’s nose and ear passages before the problem bacteria can rebound. And Israeli pediatricians, in turn, report they have reduced the rate of diarrheal infections in daycare infants by giving them formula fortified with the kinds of beneficial bacteria found in yogurt and cultured milk.
How much farther might we take such an approach? With the guiding principle that nature abhors a vacuum, a small but growing number of scientists are proposing that the only truly long-term solutions to beating disease-causing bacteria lie in promoting healthful ones.
“To look at it another way,” says Salyers, “each of us is a microbial planet, for good or for bad.” As on another planet currently under siege, the possibility exists that making peace may ultimately prove more fruitful than waging war.
Jessica, a freelance writer in Maplewood, NJ, is working on a book exploring our symbiotic relationship with the microbes that imbue our bodies. Its working title is Living with Microbes: Health and Survival in a Bacterial World, and will be published by FSG next year.
©2005 Jessica Snyder Sachs.
